|
Model 1 ("null"): BinomialLogistic( 2000 samples ): (outcome=1) ~ baseline/outcome=1 ( 100 snps) "parasite.bgen (bgen v1.2 2000 named samples zlib compression)" ( 100 snps) "host.bgen (bgen v1.2 2000 named samples zlib compression)" A basic run looks like this: $ hptest_v2.1-dev -outcome parasite.bgen -predictor host.bgen -s samples.sample -o test.csv HPTEST prints a wealth of information to the output file (and/or to a log file specified with the -log option). a population label, with type D in the sample file.) a principal component, with type C in the sample file) or discrete (e.g. If desired, covariates can be included in the analysis using the -covariates option.By default, both a Bayes factor and an approximate P-value (computed under assumptions about the asymptotic normality of the likelihood) are output. 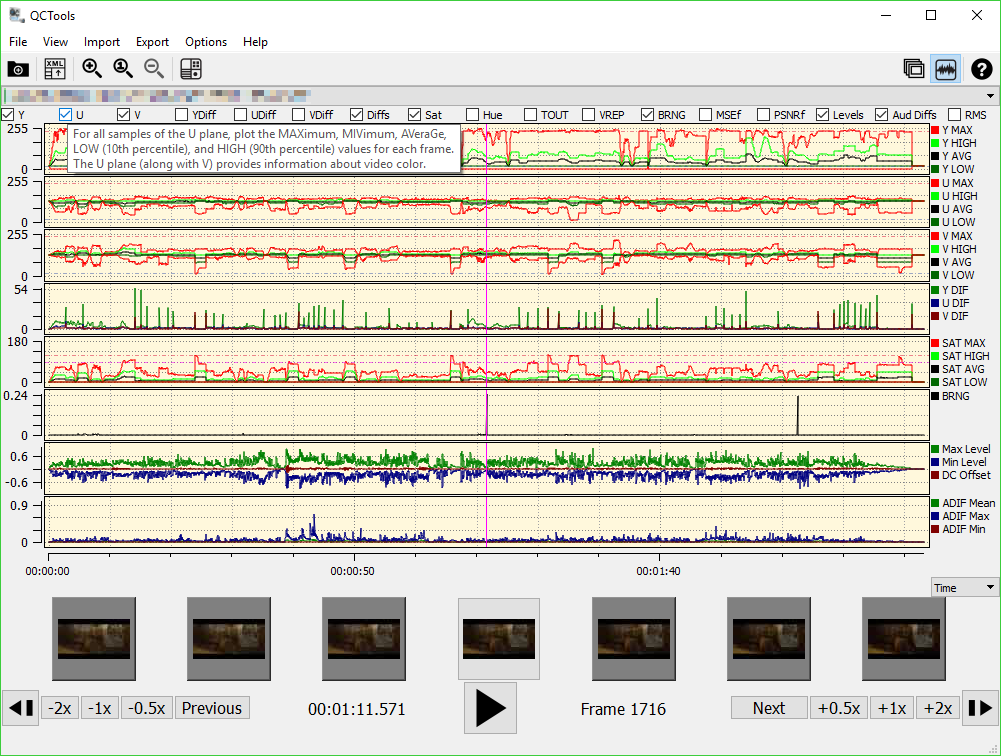
The prior can be altered using the -prior option, or turned off entirely using the -no-prior option. Specifically, a logF(2,2) prior is applied to the additive effect this is a lot like a normal distribution with mean 0 and standard deviation around 1.5, but has somewhat fatter tails. Parameters are estimated under a weak regularising prior by default.Multiple models can be specified in the same run. Alternative models can be specified using the -model option. HPTEST currently assumes predictor genotypes are diploid, and uses a 'general' model for the predictor by default, including both an additive and an overdominance term.When the outcome is diploid or higher ploidy, this is the same except that the outcome genotypes are treated as arising from multiple independent draws of an allele with the modelled probability distribution. When the outcome is haploid, this is the same as regular logistic regression. The model is binomial logistic regression, for an effect of the predictor genotype on the outcome genotype.Since rare variants are likely not very useful, by default the rest of the analysis is skipped if the count of the minor allele is < 10 in either predictor or outcome genotypes. In the latter case, for outcome genotypes a hard call will be made using a genotype probability threshold set by the -outcome-genotype-call-threshold option (default 0.9). GP field in a vcf file, or a BGEN format file). GT field in a vcf file), or else can be imputed genotype probabilities (e.g. The -reorder option to QCTOOL can help to set this up.įor each predictor and each outcome variant this does the following: All three files must represent the same set of individuals and they must be in the same order in each file. Where outcome.vcf and host.vcf contain genotype calls, and samples.sample contains sample information. In the most basic form hptest is run like this: hptest -outcome outcome.vcf -predictor predictor.vcf -s samples.sample -o test.csv Source code for HPTEST can be found here. Download and compile QCTOOL according to the compilation instructions if all goes well then the compiled application, called hptest- will appear in build/release/. 
To get HPTEST you need the development branch (named default). HPTEST is currently included as part of the QCTOOL package: HPTEST is still at a very early stage of development - if you use it, please treat with caution, and carefully sanity check any results. incl-samples, -excl-samples-where, or -incl-outcome-range - see the output of hptest -help for more). Additional options allow you to add covariates ( -covariates), to adjust the prior being used ( -prior), or to filter the list of samples or variants included (e.g. The genotype counts, maximum posterior estimates and other quantities are output to the output file ( test.csv). Regresses each parasite variant (from parasite.vcf) on each host variant (from host.vcf). Hptest -outcome parasite.vcf -predictor host.vcf -s samples.sample -o test.csv By default this includes an additional regularising prior. HPTEST finds the parameters maximising the corresponding likelihood. In which beta is a vector of parameters that is to be optimised over (the regression coefficients). Logodds( outcome genotype ) = beta * ( predictor genotype, covariates.) To do this, it implements a binomial logistic regression model: Hptest is a program for testing correlations between two sets of genotypes - for example, between genotypes of a host and of parasites infecting those hosts.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
